SHADE: A Multilevel Bayesian Framework for Modeling Directional Spatial Interactions in Tissue Microenvironments
Summary
Understanding spatial interactions between cell types is essential for interpreting immune function, tumor progression, and tissue organization. While conventional spatial statistics often assume symmetric interactions and analyze images independently, biological systems like the tumor microenvironment frequently exhibit asymmetric, hierarchically structured spatial patterns. We introduce SHADE (Spatial Hierarchical Asymmetry via Directional Estimation), a Bayesian modeling framework that captures direction-specific cell-cell associations across multiple spatial scales using smooth interaction curves and multilevel inference.
Workflow
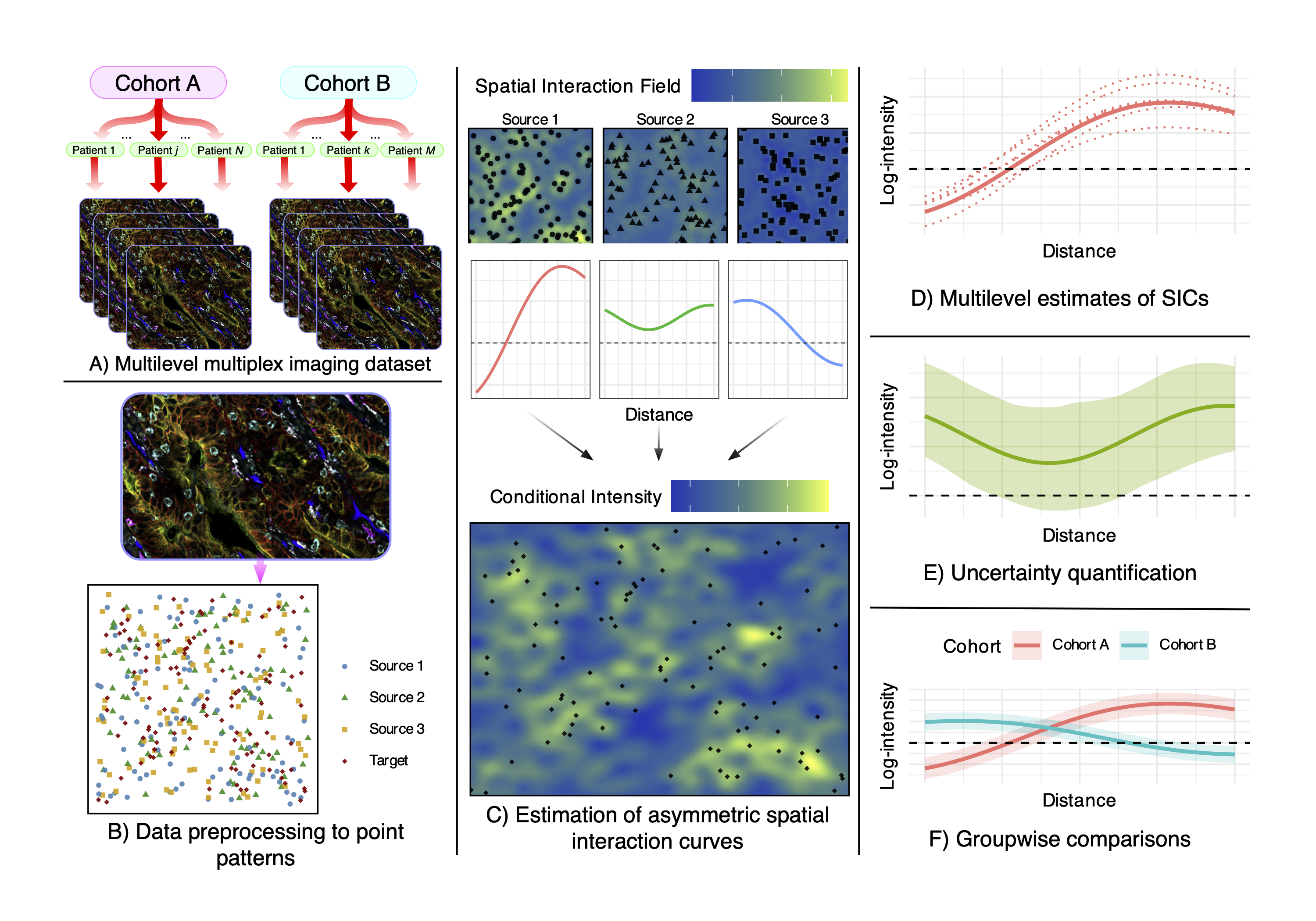
The workflow begins with hierarchically structured image data (cohorts → patients → images), which are processed into annotated spatial point patterns. SHADE then estimates Spatial Interaction Curves (SICs) to capture direction-specific cell-cell associations at multiple biological levels, allowing for analysis of spatial heterogeneity and uncertainty. These estimates can be compared across cohorts to identify differences in spatial organization.
Key Results
- Outperforms standard methods: In simulation studies, SHADE’s hierarchical pooling achieves substantially higher detection power than G-cross and K-cross envelope tests, especially under low cell densities where traditional methods struggle.
- Colorectal cancer application: Applied to 35 patients (140 images) from a multiplexed imaging dataset, SHADE identified directional spatial patterns between immune and stromal cell types while controlling for tissue architecture confounders via compartment-adjusted models.
- Multiscale heterogeneity: SHADE decomposes spatial variability across image, patient, and cohort levels, revealing that local microenvironmental structure varies considerably across patients even within the same molecular subtype.
- Scalable: Computational benchmarks show tractable runtimes even for large datasets (100K cells in ~36s, 250K cells in ~133s via variational inference).
Links
- Paper: SHADE: A multilevel Bayesian framework for modeling directional spatial interactions in tissue microenvironments — PLOS Computational Biology, 2026
- R package
- Paper reproduction code
- Vignette